This blog originated on the NIH Intramural Research Program’s “I Am Intramural” blog. Thanks to them for allowing us to repost it here.
First-Trimester Blood Analysis Could Enable Earlier, More Effective Intervention
Imagine a world in which pregnant women routinely travel to places of healing and meet with wise sages who examine a bit of their blood to divine when their babies will be born. While this may sound like something out of Greek mythology, it may soon become a reality, as researchers from the NIH Intramural Research Program (IRP) have developed a test that was able to use blood samples taken early in pregnancy to identify women who would later deliver their babies prematurely.
 |
By analyzing blood samples taken in the first trimester of pregnancy, IRP researchers were able to accurately identify most of the women who later went on to deliver their babies prematurely.
|
Women typically expect to deliver their babies after carrying them for roughly 40 weeks, a period of time known as gestation. In reality, only four percent of mothers deliver on their ‘due date.’ Roughly one in ten babies is born ‘premature’ or ‘preterm,’ meaning they are delivered before 37 weeks of gestation, and most of these births are ‘spontaneous,’ occuring with no prior warning. The earlier in pregnancy a baby is born, the higher the baby’s odds of experiencing short- and long-term health effects, including behavioral problems, learning disabilities, breathing difficulties, and infections. What’s more, because African American women deliver preterm much more often than other groups of women, premature birth is a significant contributor to racial health disparities.
If doctors were able to predict preterm births, they could intervene early to delay delivery. Right now, the best predictors for premature births are a history of preterm delivery and a shorter-than-normal cervix, the tube of tissue connecting the vagina and uterus. However, the former factor cannot be applied to first-time mothers, and most premature births occur in women who have neither of those risk factors. Measuring cervical length also requires significant resources and is only done 16 to 24 weeks into pregnancy.
 |
Senior author Dr. Roberto Romero
|
“Cervical length is the most powerful predictor of spontaneous preterm birth, but the patient must be seen at a healthcare facility and we need equipment and expertise,” says IRP senior investigator Roberto Romero, M.D., D.Med.Sci., the new study’s senior author. “It would be great if we could have a simple blood test that could predict spontaneous preterm delivery.”
The IRP team’s study used blood samples collected from women who had been pregnant for 6 to 13 weeks, significantly earlier than cervical length can be used to predict preterm birth. First, the researchers analyzed samples from nine women who went on to deliver preterm and 70 women who delivered at term to measure the concentrations in their blood of 45 different microRNA molecules — short strands of genetic material that help control the behavior of genes. This analysis showed that 12 specific microRNAs were much more abundant in blood from women who delivered prematurely.
Next, using blood collected from 78 other women during the same early period of pregnancy, Dr. Romero’s team measured the levels of those 12 microRNAs to attempt to retroactively identify which of them ultimately delivered prematurely. Overall, their analysis method correctly classified nearly 90 percent of the women who went on to deliver preterm, with a low rate of false negatives. Moreover, when the IRP researchers separately analyzed samples from women whose blood was taken before 10 weeks of gestation and those whose blood was taken after that time point, they found that they were able to identify 80 percent of the women who went on to deliver prematurely in the former group and all of the women who delivered preterm in the latter group.
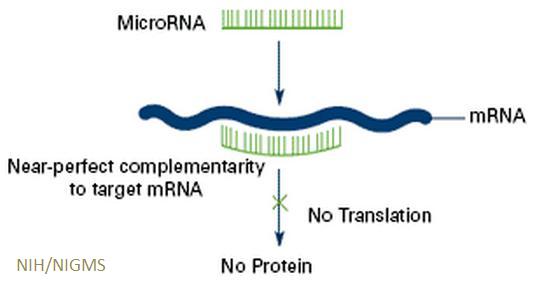 |
Diagram of how micoRNAs affect protein production by genes
|
MicroRNAs regulate the activity of genes by reducing the amount of protein they produce.
“I never want to claim anything is 100 percent, but the point is if we have a blood test based on microRNAs, we can predict spontaneous preterm delivery,” Dr. Romero says. “The idea that there is a blood test that can help us assess a person’s risk for preterm delivery is not a dream.”
The researchers also consulted an online database to identify biological pathways that those 12 microRNAs are involved in. These include processes that help transport biological molecules around the cell, control the activity of estrogen-related genes, and activate enzymes involved in cell death. This information provides clues as to the biological triggers of some cases of preterm birth, knowledge that will help scientists develop therapies that prevent it.
Meanwhile, doctors already have some methods to reduce the risk of preterm birth, and these approaches are more effective the earlier in pregnancy they are implemented. If future studies in larger and more diverse populations confirm the results of Dr. Romero’s research, clinicians could identify patients who may need those treatments using a simple blood test.
“We are interested not just in prediction but also in prevention,” Dr. Romero says. “There are substantial advantages if we can use blood markers during pregnancy for this purpose, and the earlier we can do that the better because then there is a window in which we can intervene.”
Reference
Winger EE, Reed JL, Ji X, Gomez-Lopez N, Pacora P, Romero R. MicroRNAs isolated from peripheral blood in the first trimester predict spontaneous preterm birth. (2020) PLoS ONE 15:e0236805. doi: 10.1371/journal.pone.0236805. PMID: 32790689.