1-UG3-TR002884-01 • The Ohio State University
1-UG3-TR002884-01 • The Ohio State University
Principal Investigator:
Contributors: L. James Lee, Inyoul Lee, Kai Wang, Betty YS Kim
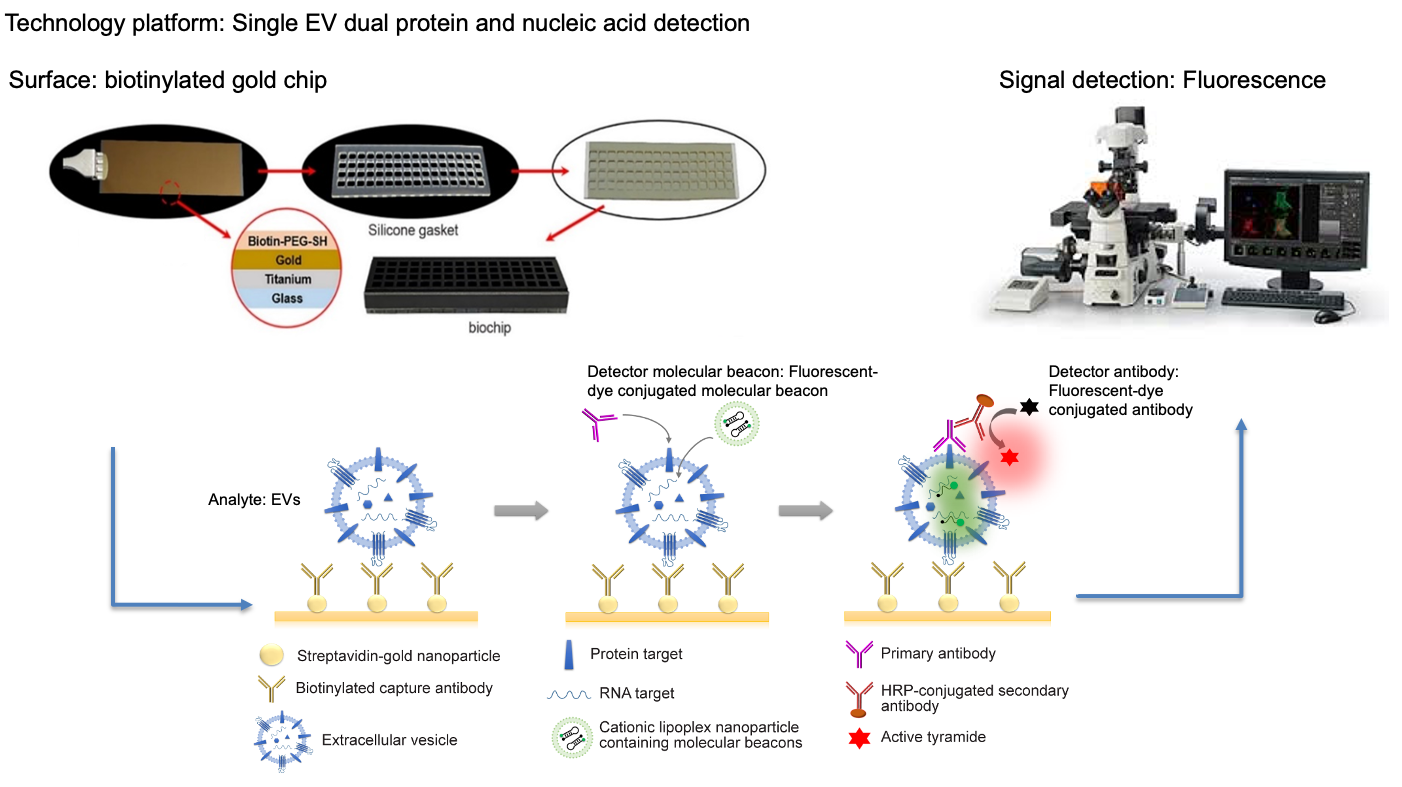
Extracellular vesicles (EVs) such as microvesicles and exosomes are small membrane vesicles released by cells in the body. EVs are present in all biological fluids tested (e.g., blood, urine, cerebral spinal fluid) and contain various biomolecules including DNAs, RNAs, proteins and metabolites, and have been implicated as part of the cell-cell communication systems. Despite their importance, the current methods of isolating and characterizing EVs are technically challenging. The isolation methods usually cumbersome and irreproducible, and the characterization relies on techniques like Polymerase Chain Reaction (PCR), Next Generation Sequencing (NGS), and Mass Spectroscopy (MS) which just provide an aggregate of the overall RNA/DNA and protein content. During the characterization process, EVs are broken down to obtain their internal contents. Consequently, the molecular information at individual EV is lost. Given the heterogeneity of EVs, it is imperative to study EV-mediated intercellular signaling processes at the single EV level in order to gain important insights of their effects on promoting drug resistance, immunosuppression, epithelial-to- mesenchymal transition (EMT), cancer metastasis, and cachexia. Therefore, there is a critical need to develop technologies that provide accurate and efficient analysis of the molecular content within individual EVs. We propose an integrated system using a size exclusion chromatography to first sort the EVs in biofluids into well- defined size-based subpopulations, and then distribute each subpopulation into a set of parallel microfluidic channels where each channel is patterned with a number of microdomains tethered with antibody-conjugated liposomal nanoparticles containing molecular beacons (MBs) for enriched isolating/capturing of specific membrane protein/peptide-rich single EVs in the subpopulation, and simultaneously identification of specific RNA targets via MB-RNA hybridization when the captured EVs are fused with liposomes. Fluorescence- labelled antibodies may also be added to each microchannel to quantify the target membrane protein content of the captured single EVs. The development and feasibility demonstration of this novel technology will be conducted in the UG3 phase with a small-scale array for selected RNA and protein targets using well-characterized synthetic vesicles (SVs), EVs released from glioblastoma (GBM) cell lines, and spiked EVs in normal donor serum. In the UH3 phase, we plan to scale-up the biochip system design for high-throughput using the GMP type biochip fabrication. The applicability of this new technology will be validated for EVs from GBM cell lines as well as serum and cerebral spinal fluid (CSF) from GBM patients. In both phases, EV-based cell-cell communication will be investigated to determine if and how specific GBM EV subpopulations are involved in immune-regulation within GBMs. We have assembled a multi-disciplinary team with extensive knowledge and experience in nanobiotechnology, microfluidics, EV characterization, micro/nano-fabrication, EV RNA profiling and biomarker discovery, GBM diagnosis and treatment, and biostatistical analysis. The proposed aims and milestones are given as follows: UG3 Phase- Specific Aim 1: Development of a biochip to capture and characterize specific EV subpopulations at single EV level. Specific Aim 2: Comparison of results between the single EV-based measurements and conventional total EV-based averaged measurements. Quantitative Milestones: (i) Sorting, isolation and quantitative analysis of selected mRNA and miRNA targets in single SVs and EVs with >90% repeatability and better EV enrichment than conventional ultracentrifugation and antibody-based microfluidics methods; (ii) Identifying one or more EV subpopulations for high sensitivity detection of GBM cell- derived EVs; (iii) Identifying one or more GBM EV subpopulations which may involve in immuno-regulation. UH3 Phase- Specific Aim 1: Scaleup of the biochip manufacturing. Specific Aim 2: To perform EV analysis on clinical samples from GBM patients. Quantitative Milestones: (i) Sorting, isolation and quantitative analysis of mRNA/miRNA and membrane protein targets in single EVs from both blood and CSF samples with >90% repeatability and 5-fold better EV enrichment than conventional ultracentrifugation and antibody-based microfluidics methods; (ii) <10% false positive/negative prediction from a total of 120 GBM patients and non-patient samples; (iii) Identifying one or more GBM EV subpopulations which may involve in immunosuppression and/or associated with worse clinical outcomes.
Related Projects