When someone finds out that I work at the National Institute on Aging, they usually ask me “How can I stop aging?”. Although aging is inevitable, it is well-known that individuals age at different rates and that certain groups age more rapidly than others. Hence, one of our laboratory’s objectives is to identify why certain population groups age differently, with the goal to find biomarkers that can tell us an individual’s biological age (how old you seem) versus chronological age (the actual time you have been alive).
Several years ago, we began this journey by examining whether small non-coding RNAs called microRNAs (miRNAs) change with age. We focused first on these regulatory RNAs because data has shown that these RNAs are particularly stable in biofluids, such as serum and plasma, and can be identified readily using small RNA sequencing. We identified several serum miRNAs that change significantly with age in both humans and rhesus monkeys (Noren Hooten, Fitzpatrick, et al. 2013). In most cases, their abundance in circulation decreases as we get older. Interestingly, some of these miRNAs target and negatively regulate the expression of various inflammatory markers, suggesting that decreased expression of these circulating miRNAs may contribute to higher levels of inflammatory markers that have already been observed in the elderly.
More recently we have focused on establishing a more complete extracellular RNA (exRNA) profile of human aging. To do so, we developed a sequencing pipeline that enables us to sequence both small and long RNAs in one sequencing reaction, which lowers both the cost and labor required (Dluzen, Noren Hooten, et al. 2018). Cataloging what is the “normal” distribution of exRNA in young and old individuals and identifying age-dependent differences will aid in establishing important references for the study of age-related disease. The Ensembl database classifies RNA into various categories, termed biotypes. We found that most RNA biotypes were similar in distribution between young and old, but several biotypes, including mitochondrial tRNAs (Mt_tRNA), mitochondrial ribosomal RNAs (Mt_rRNA), and unprocessed pseudogenes, were significantly higher in older individuals.
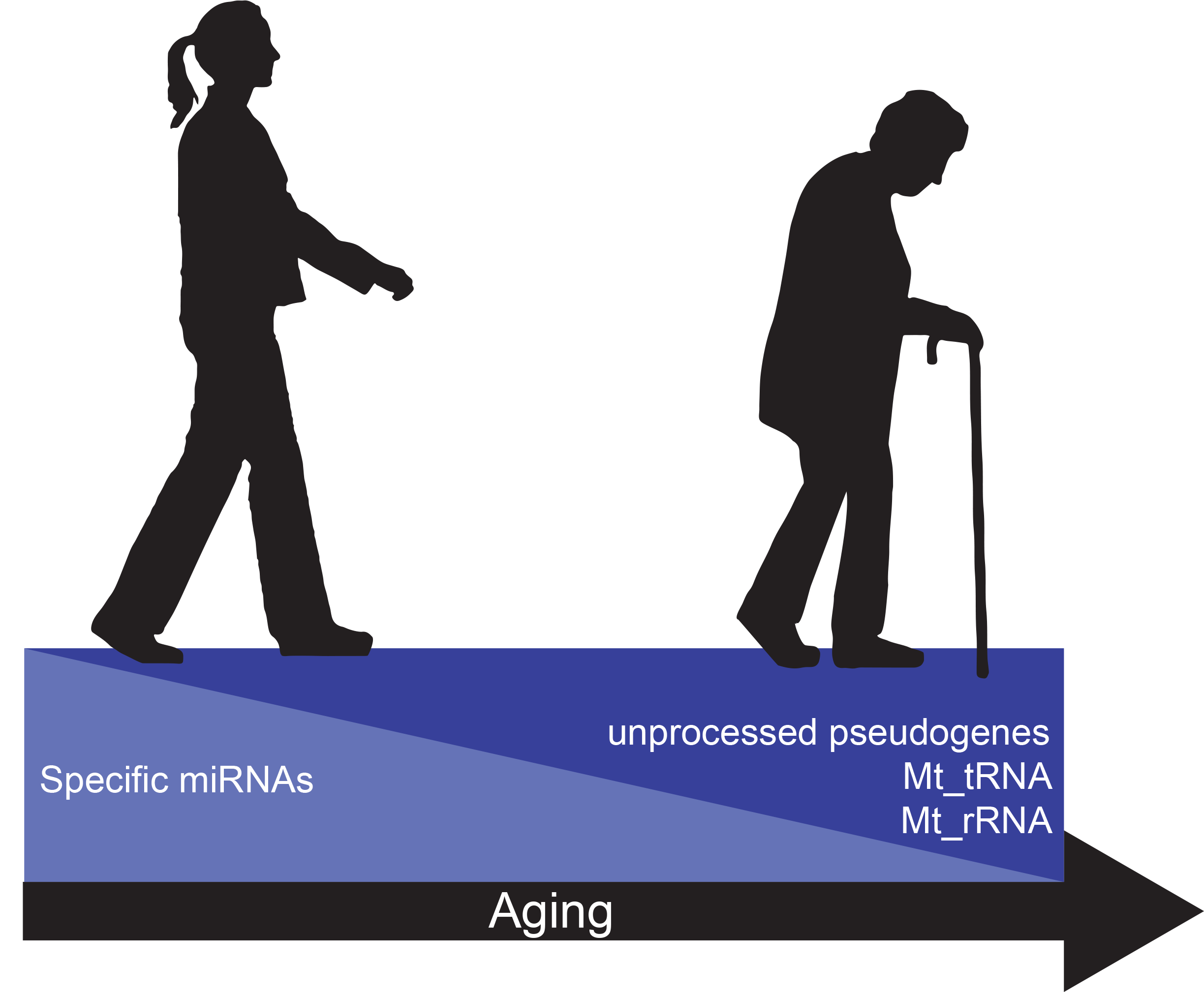
Figure 1. Changes in circulating factors with age. A decrease in circulating levels of specific miRNAs occurs with age. On the other hand, unprocessed pseudogenes, Mt_tRNAs, and Mt_rRNAs increase in abundance with age.
Pathway analysis revealed that RNAs related to mitochondria, response to oxidative stress, and chromatin remodeling were all enriched in the circulation of older individuals, providing potential clues as to what pathways may be deregulated as humans age. We also further validated our sequencing results in a larger cohort of individuals and found age-related changes in a messenger RNA, a small nucleolar RNA, a pseudogene transcript, a small nuclear pseudogene transcript, and several additional miRNAs.
What was very interesting was that we identified many circular RNAs (circRNAs) in serum, which we have named ex-circRNAs. Recent attention has been focused on circRNAs, as this class of ncRNAs may be important modulators of gene expression. CircRNAs have long half-lives (i.e. are stable) compared to mRNAs, making them an attractive new serum biomarker. However, little is known about ex-circRNAs, and I anticipate that this will be an active area of interest in the coming years.
As we have begun to establish an exRNA profile of human aging, there remain important unanswered questions in the field. Currently, we do not fully understand how exRNA in the circulation reflects the health status of our cells and tissues. It has also proven difficult to ascertain which cell type is contributing exRNA into the circulation. Further research is needed to better understand these questions.
Our identification of changes in circulating miRNAs and exRNAs establishes baseline references for how these biomarkers change with human age. As the risk for many diseases including cancer, heart disease, and neurological diseases increase with age, it is important to consider age when examining these factors in relation to a specific disease. Although we have not identified the “fountain of youth” or the “magic elixir” for aging, we hope that establishing these profiles with normal aging will soon help to identify circulating biomarkers that can distinguish individuals with faster biological aging that may result in shortened health span and life span.
References
Dluzen DF, Noren Hooten N, De S, Wood H, Zhang Y, Becker KG, Zonderman AB, Tanaka T, Ferrucci L & Evans MK. Extracellular RNA profiles with human age. Aging Cell 2018;e12785. PMID 29797538.
Noren Hooten N, Fitzpatrick M, Wood WH, De S, Ejiogu N, Zhang Y, Mattison JA, Becker JG, Zonderman AB & Evans MK. Age-related changes in microRNA levels in serum. Aging (2013) 5: 725-740. PMID 24088671.