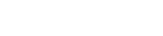|
This blog originated as a press release from the Johns Hopkins Kimmel Cancer Center. Thanks to them for allowing us to repost it here. |
It may be possible to identify the presence of an aggressive brain tumor in children by studying their cerebrospinal fluid, according to new research led by Johns Hopkins Kimmel Cancer Center investigators.
Comparing cerebrospinal fluid samples from 40 patients with medulloblastoma — the most common malignant brain tumor in children, accounting for 10% to 15% of pediatric central nervous system tumors — and from 11 healthy children without the disease, investigators identified 110 genes, 10 types of RNA–the machinery that translates proteins–called circular RNAs, 14 lipids and several metabolites that were expressed differently between the two groups. While these details were not specific enough to distinguish among the four subtypes of medulloblastoma, they could be used to identify the presence of cancer versus normal fluid. A description of the work was published in the journal Acta Neuropathologica Communications.