Heart failure and sudden cardiac death are end-stage manifestations of coronary heart disease worldwide. With an aging population and improvements in therapy for coronary revascularization during myocardial infarction, the number of patients proceeding to advanced heart failure is growing. The role of extracellular RNAs (exRNAs) in the field of cardiovascular medicine has been growing in parallel, as diagnostic, prognostic, and potentially even mechanistically and physiologically relevant biomarkers.
Our interest in this field grew out of an initial foray into the world of advanced heart failure patients referred for biventricular pacemaker therapy. Ventricular “dyssynchrony” due to delayed activation of the left ventricular (LV) lateral wall is present in nearly 50% of patients with symptomatic advanced heart failure (HF) and reduces effective LV function. Cardiac resynchronization therapy (CRT) is a type of pacemaker that can reduce dyssynchrony and mitigates adverse cardiac remodeling and improves prognosis in up to 65-70% of individuals. Unfortunately, even in patients who meet criteria for CRT, 30% or more do not derive hemodynamic or clinical benefit, and efforts to define clinical, image-based, plasma or electrocardiographic biomarkers to predict responsive patients have not been very successful. Given the role of miRNAs in regulating gene networks relevant to cardiovascular diseases (e.g., fibrosis, arrhythmia), we became interested in investigating ex-RNAs as potential markers of CRT responsiveness.
To identify circulating miRNAs that predict response to CRT, we assessed plasma miRNA profiles prior to CRT in patients with advanced HF and dyssynchrony (HFDYS) with or without subsequent echocardiographic improvement after CRT. In this group, we discovered a set of miRNAs disregulated in responders, but focused after statistical efforts on microRNA-30d, which was a strong, independent marker of risk in adjusted logistic and linear regression. Baseline concentrations of miRNA-30d were associated with CRT response.
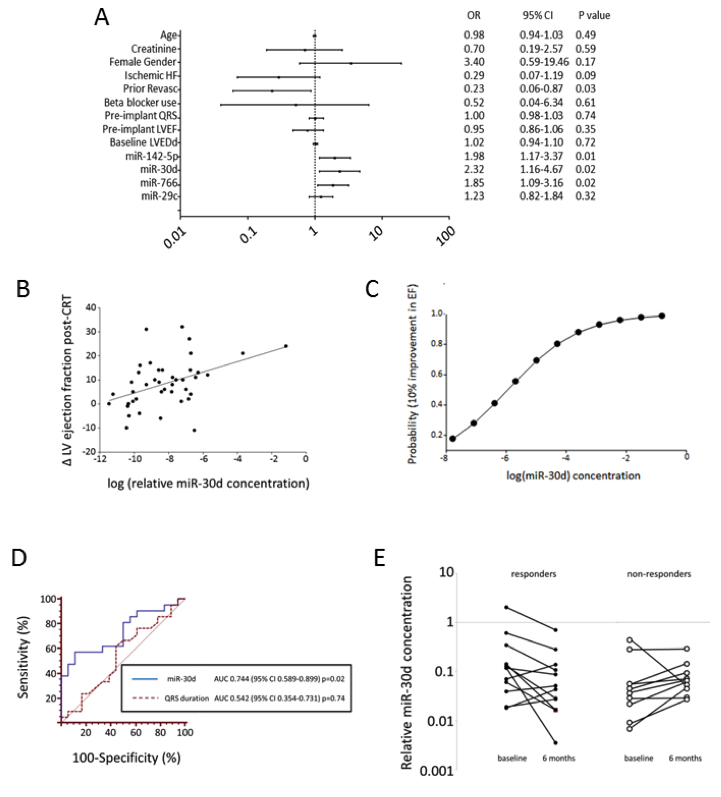
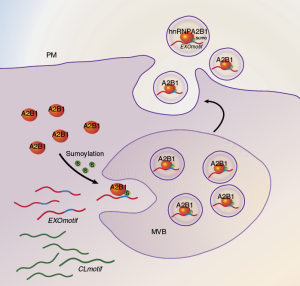
Clinical research identifies a novel ex-RNA biomarker of disease. (A) Logistic regression for CRT responsiveness; (B) Linear regression of changes in LV ejection fraction post-CRT with miR-30d concentration; (C) Logistic probability function of responsiveness by miR-30d; (D) ROC curve of miR-30d versus QRS duration. (E) Changes in miR-30d at 6 months after CRT.
Given that CRT responsiveness may derive from “fixing” the electrical delays between the lateral and septal wall of the ventricle, we then found that miR-30d is more highly expressed in the lateral wall of the LV in a canine model of dyssynchrony. Mechanistically, miR-30d was synthesized and released by cardiomyocytes (CM) in response to increased mechanical stress, and mediated CM hypertrophy with adaptive features, including protection against TNF-α-induced CM apoptosis.
To further explore the functional role of miR-30d, we identified miR-30d targets that are dynamically regulated in the canine model of HFDYS. Using a LIMMA (linear models for microarray data) approach, genes whose expression was inversely correlated to miR-30d expression (i.e. genes which were down-regulated in the lateral wall of dyssynchronous dogs where miR-30d levels were highest compared to the septal wall) were identified, and we screened these and other putative miR-30d targets using the Tarbase and TargetScan prediction algorithms. Subsequently, using the IPA (Ingenuity Pathways Analysis) approach we generated predicted functional pathways associated with the observed changes in miR-30d expression. Several of these genes affect multiple biologically relevant pathways in the heart, specifically, LIMS1, PPP1R14c, MAK3K13, and JAK1. In addition, several other miR-30d targets with known roles in cardiac hypertrophy were identified using Tarbase. Intriguingly, one of these, mitogen associated protein kinase 4 (MAP4K4), a downstream mediator of tumor necrosis factor-alpha (TNF-α) has recently been identified as a target of miR-30d in pancreatic tissue 1 and has been shown to play a key role in TNF-mediated inflammatory processes. Because TNF-α not only plays an important role in pathogenesis of cardiac remodeling, but also is differentially expressed in the lateral and septal walls in the HFDYS canine model 2, we deemed MAP4K4 a potentially important target of miR-30d in HFDYS.
To validate candidates identified through bioinformatic analyses, we assessed the ability of miR-30d overexpression in CMs to effectively silence the mRNAs for the putative targets. Transient transfection of CMs with miR-30d caused a significant decrease in the mRNA levels of LIMS1, MAP4K4, and PPP1R14c relative to scramble transfected cells. Given the involvement of TNF-α signaling in the pathophysiology of HFDYS as described above, we focused on the interaction between miR-30d and MAP4K4. MAP4K4 protein was significantly decreased in the lateral wall in HFDYS compared to lateral wall of control animals, correlating inversely with miR-30d levels in these two regions. Conversely, in the CRT lateral wall (where wall stress presumably decreased with resynchronization), miR-30d levels fell compared to HFDYS, and MAP4K4 protein level returned to control levels, suggesting that MAP4K4 was an in vivo target of miR-30d in these models. Treatment of scramble transfected CMs with TNF-α led to a robust increase in MAP4K4 mRNA, which was markedly attenuated by miR-30d overexpression. miR-30d transfection inhibited TNF-α mediated apoptosis in CMs, suggesting that miR-30d may be protective against the maladaptive effects of TNF-α. miR-30d transfection also prevented TNF-α-mediated increase in molecular markers associated with pathological hypertrophy and fibrosis (ANP, TIMP1 and β-MHC).
Finally, we measured levels of high sensitivity troponin T, a marker of myocardial injury, in our patients, demonstrating an inverse association between miR-30d and troponin TnT (Spearman r=-0.51, p=0.001), suggesting that higher levels of miR-30d may be cardioprotective in human HF patients.
These experiments provide ‘in vivo’ supportive evidence for our hypothesis that miR-30d may be protective against inflammation (TNF-α) induced cardiomyocyte injury, thereby promoting cardiomyocyte survival and favorably influencing the degree of cardiac remodeling in response to CRT. These results support the hypothesis that clinically useful ex-RNA biomarkers may be functionally implicated in disease pathogenesis.
As part of the exRNA consortium, our group hopes to extend these results to a population of patients at risk for LV remodeling (post-MI), to determine whether this approach (biomarker discovery through clinical research; validation through basic research; and reapplication for disease diagnostics and physiology through translational research) is feasible in developing novel biomarkers of human cardiovascular disease.