A recent study by Wei et al., 2017 catalogs the composition and characteristics of extracellular RNA (exRNA) secreted via three different routes from parent cells. The work provides novel insights into the biology of exRNA transport and intercellular communication, as well as the clinical potential of exRNA as a biomarker of disease.
The senior investigator of the study, Anna Krichevsky, Ph.D., at Brigham and Women’s Hospital and Harvard Medical School described the rationale for the study: “To understand the functions of exRNA complexes, we first have to define the exRNA repertoire with minimal bias.”
The exRNA Composition of Microvesicles, Exosomes, and Ribonucleoproteins (RNPs)
Dr. Krichevsky’s group sequenced RNA in microvesicles, exosomes, and extravesicular ribonucleoproteins (RNPs) isolated from glioma stem cells. They found that the majority of exRNA in all three fractions is noncoding. Although most exRNA studies focus on one class of noncoding RNA called microRNA (miRNA; 21-25 nucleotide molecules that repress gene expression), they reported that <10% of exRNA secreted by glioma stem cells is miRNA.
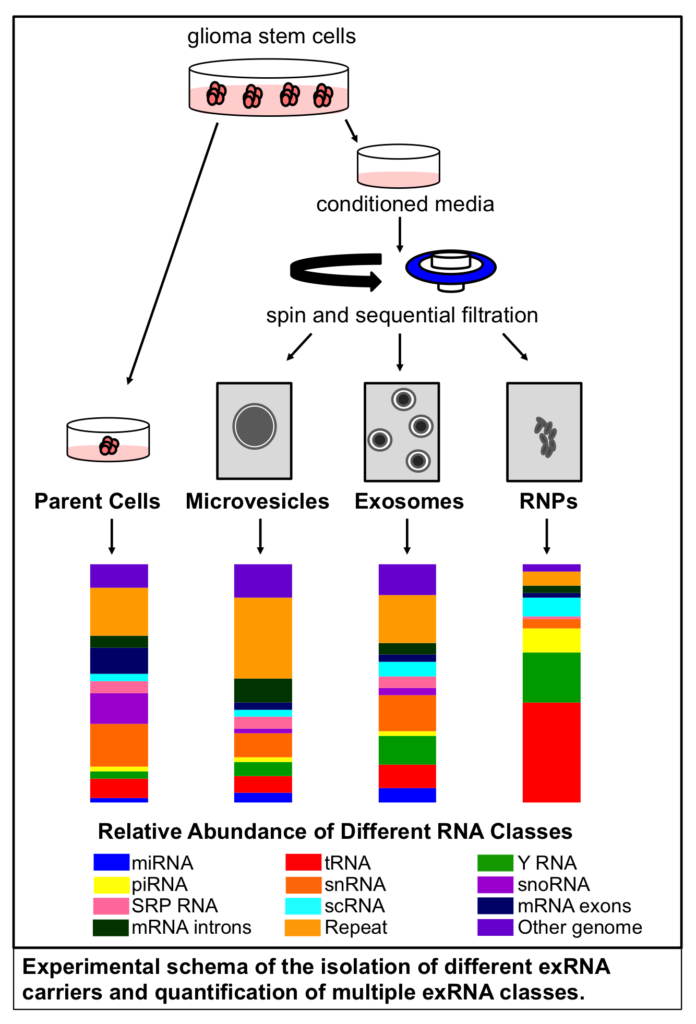
Comparing the profile of exRNA isolated from RNPs and extracellular vesicles (EVs) — including both exosomes and microvesicles, they found that RNPs contain higher amounts of noncoding cytoplasmic Y RNA and transfer RNA (tRNA) fragments.
Y RNA folds into a characteristic stem-loop structure and was originally found in protein-RNA complexes of individuals with autoimmune diseases. According to Dr. Krichevsky, “Despite abundant expression in all vertebrate cells, the physiological functions of Y RNA are only beginning to emerge.” Some evidence suggests that Y RNA plays a role in DNA replication and RNA quality control. Y RNA fragments may also be involved in cell death, ribosomal RNA maintenance, and histone gene expression.
tRNA is well known as a mediator of the translation of mRNA to protein. However, recent studies suggest that tRNA fragments found in exRNA are involved in regulating gene expression during cellular stress responses. Dr. Krichevsky discussed the implications of extracellular tRNA on cellular communication: “Based on the exRNA levels and biological functions of tRNA, we hypothesize that transferred tRNA transcripts can have a major impact on recipient cells.”
Along with noncoding RNA, a small proportion of gene-encoding messenger RNA (mRNA) was detected in extracellular vesicles and RNPs. Previous studies have also found extracellular mRNA; however, they did not determine if the mRNA transcripts were intact or fragmented. Dr. Krichevsky’s group was the first to show that short (<1000 nucleotides), endogenous full-length mRNAs can be packaged into exosomes, and microvesicles contain even longer mRNAs. Fragments of long mRNA transcripts were also present in exRNA.
Using exRNA as a Biomarker
Researchers are currently exploring the use of exRNAs as potential biomarkers for the diagnosis and monitoring of diseases. However, many exRNA biomarker studies have been limited in scope because they examined a heterogeneous pool of exRNA purified from an unfractionated collection of EV types.
By fractionating conditioned media from glioma stem cells into microvesicle, exosome, and RNP fractions, Dr. Krichevsky’s group was able to compare their RNA profiles with those of the parent cells. They found that the RNA content of microvesicles most closely resembled that of the parent cells, making this type of exRNA carrier a good candidate for disease biomarkers.
Dr. Krichevsky said, “We believe there is more intact mRNA in microvesicles, which we can consider for biomarkers. We can think about genes that are mutated. On the other hand, miRNAs are more enriched in the exosomes. It would be great if we could detect cancer mutations and non-coding RNA biomarkers in biofluids; then we would not need to do a biopsy.”
Conclusions
By developing novel experimental approaches to illustrate that exRNA composition differs by exRNA carrier, Krichevsky’s group has made significant contributions to exRNA research. Moreover, they highlighted that exRNA contains more than the well-studied miRNAs, including full-length mRNA molecules, Y RNA, and tRNA. Their data also indicate that the RNA profile of microvesicles is most similar to that of the cell of origin, including the presence of full-length mRNAs, making microvesicle exRNA a good candidate for some disease biomarkers.
In an interview, Dr. Krichevsky discussed the importance of this study: “Our work changed the way people thought about exRNA by showing them the exRNA in numbers, which helps appreciate their heterogeneity and the overall impact. The field is shifting from focusing on a specific extracellular miRNA to now considering that there are thousands of different RNAs present in extracellular complexes.”
References
Wei, Z. et al. Coding and noncoding landscape of extracellular RNA released by human glioma stem cells. Nature Communications (2017) 8:1145. doi:10.1038/s41467-017-01196-x
Researchers invent novel RNA nanotech to decorate exosomes for effective cancer therapy
Optimizing the production of extracellular vesicles for therapeutic applications
There are no comments.