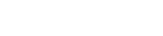This blog post originated as a press release from the University of Alabama at Birmingham.
University of Alabama at Birmingham researchers have found a novel, previously unreported pathogenic entity that is a fundamental link between chronic inflammation and tissue destruction in the lungs of patients with chronic obstructive pulmonary disease, or COPD. COPD is the fourth-leading cause of death in the world.
This pathogenic entity — exosomes from activated polymorphonuclear leukocytes, or PMNs — caused COPD damage when the small, subcellular particles, collected from purified PMNs, were instilled into the lungs of healthy mice. Remarkably, the UAB researchers also collected exosomes from the lung fluids of human patients with COPD and the lung fluids of neonatal ICU babies with the lung disease bronchopulmonary dysplasia; when those human-derived exosomes were instilled into the lungs of healthy mice, they also caused COPD lung damage. Damage was primarily from PMN-derived exosomes from the human lungs.
“This report seems to provide the first evidence of the capability of a defined non-infectious subcellular entity to recapitulate disease phenotype when transferred from human to mouse,” said J. Edwin Blalock, Ph.D., professor of pulmonary, allergy and critical care medicine in the UAB Department of Medicine. “I think this could be a very profound discovery. A lot of what we have found here will apply in other tissues, depending on the disease.”
Other diseases marked by immune cell inflammation and tissue destruction include heart attacks, metastatic cancer, and chronic kidney disease. The activated PMN exosomes may also contribute to lung damage in other lung diseases that have excessive PMN-driven inflammation, such as cystic fibrosis. The study is reported in the journal Cell.
“These findings highlight a novel role of the innate immune response in chronic lung diseases and could be used for the development of new diagnostics and therapeutics for COPD and possibly cystic fibrosis,” said James Kiley, Ph.D., director of the Division of Lung Diseases at the National Heart, Lung, and Blood Institute, part of the National Institutes of Health.
Background
COPD, a smoking-associated disease, is marked by PMN-driven inflammation in the lungs. Damage to the lung tissue leads to airway obstruction, shortness of breath, and respiratory failure. PMN immune cells, also known as neutrophils, are part of the body’s white blood cell defense against infections and tissue damage. They comprise 60 percent of the body’s white blood cells, or about 2.5 billion PMNs in each pint of blood. PMNs are voracious eaters of microbes or damaged human cells after activation by a signal of infection.
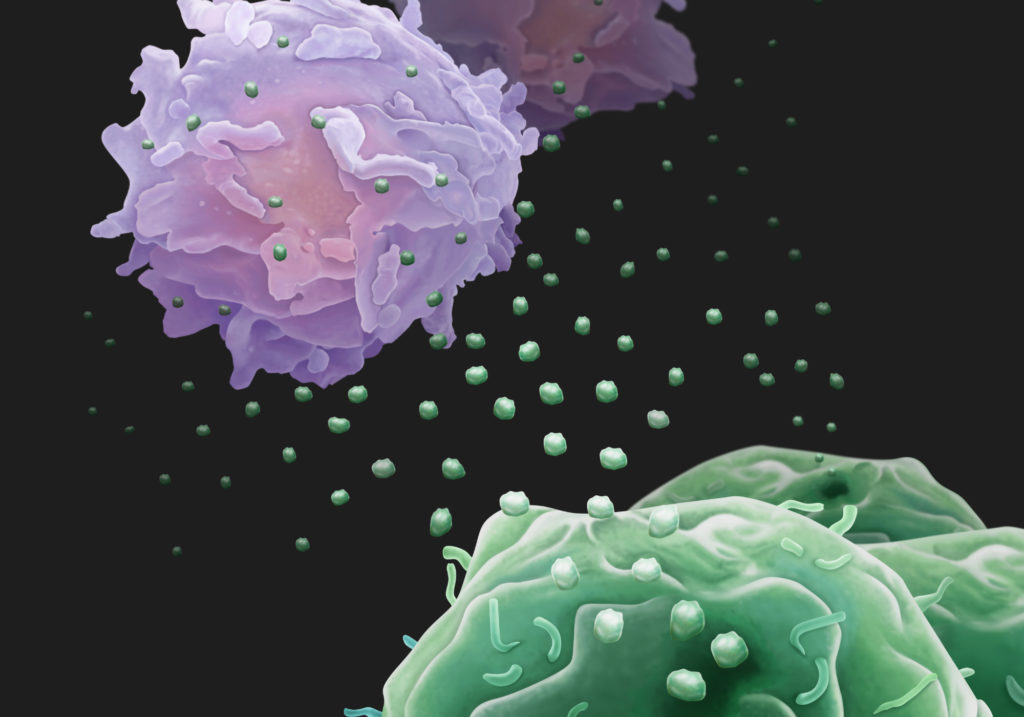
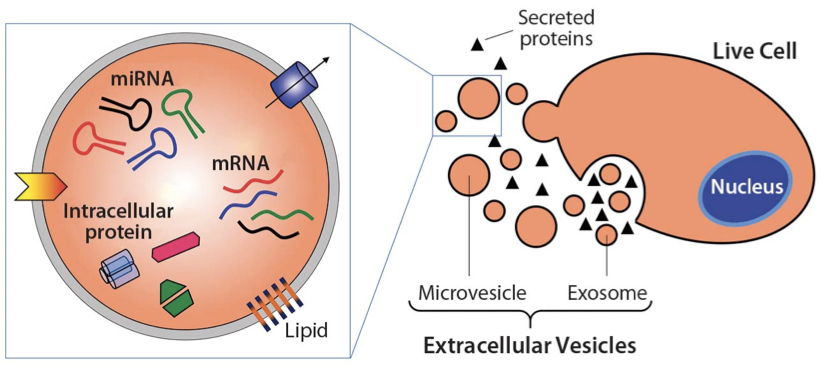
All cells shed exosomes. These tiny extracellular membrane-bound vesicles can be mediators of cell-to-cell communication, and they can ferry a diverse cargo of proteins, lipids, and nucleic acids from cell to cell. The UAB research focused on a recently found third role for exosomes — the ability to harbor protease enzymes.
Activated PMNs are known to release neutrophil elastase, or NE, a protease that can degrade type I collagen and elastin. The collagen and elastin proteins help form the extracellular matrix that glues cells together. In the lungs, the extracellular matrix and lung cells are sheets of tissue that help form the tiny alveoli, where the lung exchanges oxygen and carbon dioxide. In COPD, the damaged alveoli enlarge, reducing oxygen exchange and forcing the heart to pump harder to push blood through the lungs.
NE and other proteases from PMNs can attack microbes. Healthy lungs are protected by anti-proteases that can inhibit the proteases. Normally, NE is inhibited by a robust barrier of alpha1-antitrypsin in the lung.
The research
Blalock and fellow researchers investigated whether NE might exist in an exosomal form and whether such exosomes might bypass alpha1-antitrypsin inhibition to contribute to inflammatory lung disease.
They found that exosomes from quiescent PMNs did not cause COPD when transferred to healthy mice. In contrast, exosomes from activated PMNs did cause COPD, as measured by histologic changes of the alveoli, increased pulmonary resistance and enlargement of the right heart ventricle that pumps blood to the lung.

“This investigation reveals an entirely unappreciated aspect of the interplay between inflammation, proteolysis, and matrix remodeling with far-reaching implications for future research.”
J. Edwin Blalock
The activated PMN exosomes were covered with enzymatically active surface-bound NE, while quiescent PMN exosomes had none. This surface NE was resistant to alpha1-antitrypsin inhibition; the exosomes from activated PMNs degraded collagen, they caused emphysema when put into mouse lungs, and they carried the PMN cell-surface markers CD63 and CD66b that identify them as coming from PMNs. Human COPD lung-derived exosomes carrying those PMN cell-surface markers conferred COPD to mice.
A very large dose of purified NE — enough to overwhelm the alpha1-antitrypsin barrier — can cause alveolar enlargement in mice. Because the exosome-bound NE was protected against apha1-antitrypsin inhibition, researchers found that the dose of activated PMN exosomes needed to cause the same damage as purified NE was 10,000 times less.
The activated PMN exosomes had another cause for their aggressive proteolysis — they carried integrin Mac-1 on their surface. Integrin Mac-1 allowed the exosomes to bind directly to collagen fibrils, a second mechanism besides protected NE for why the proteolytic exosomes exert an outsized degradative capacity in relation to their size and protease load.
“This investigation reveals an entirely unappreciated aspect of the interplay between inflammation, proteolysis and matrix remodeling with far-reaching implications for future research,” Blalock said. “Our report significantly expands the biological repertoire of the exosome, demonstrating potent biological effects of these particles ex cellula.”
Looking ahead
The study also suggests therapeutic strategies to interrupt pathogenic aspects of PMN exosome function: 1) disrupting the ionic binding of the NE to the exosome, to dislodge the NE and make it susceptible to alpha1-antitrypsin; 2) inhibiting the exosomal integrin Mac-1 to block collagen binding; and 3) directly inhibiting the exosomal NE with small-molecule compounds.
Blalock is also interested in another big question — exosome activity in healthy smokers.
“Only one in seven or one in eight smokers gets COPD,” he said. “It would be an amazing outcome if we found activated PMN exosomes in a subpopulation of people who smoke.” Those people could then be warned of the risk they faced.
This Cell study took six years of work.
Significant research was done by co-first authors Kristopher Genschmer, Ph.D., and Derek W. Russell, M.D., who were NIH T32 grant trainees with Blalock. Both are assistant professors in the UAB Division of Pulmonary, Allergy and Critical Care Medicine. Amit Gaggar, M.D., Ph.D., a professor of pulmonary, allergy and critical care medicine, is co-senior author with Blalock, and he is a former trainee who did his Ph.D. with Blalock. Co-author Charitharth Vivek Lal, M.D., assistant professor in the UAB Pediatrics Division of Neonatology, is the physician who collected the lung fluid from neonates and performed all of the bronchopulmonary dysplasia work.

Co-authors with Genschmer, Russell, Gaggar, Lal and Blalock of the paper “Activated PMN exosomes: Pathogenic entities causing matrix destruction and disease in the lung” are Tomasz Szul, Mojtaba Abdul Roda, Xin Xu, Liliana Viera, Tarek H. Abdalla, Robert W. King, J. Michael Wells and Mark T. Dransfield, UAB Department of Medicine, Division of Pulmonary, Allergy and Critical Care Medicine; Preston E. Bratcher, National Jewish Medical Center, Denver, Colorado; Brett D. Noerager, University of Montevallo, Montevallo, Alabama; Gabriel Rezonzew, UAB Department of Pediatrics; Brian S. Dobosh, Camilla Margaroli and Rabindra Tirouvanziam, Department of Pediatrics, Emory University, Atlanta, Georgia; and Carmel M. McNicholas, UAB Department of Cell, Developmental and Integrative Biology.
This study was supported by National Institutes of Health grants HL135710, HL077783, HL114439, HL110950, HL126596, HL102371, HL126603, HL123940, HL105346-07 and HL105346-05; American Heart Association grant 17SDG32720009; and Veterans Affairs grant BX001756.
Blalock is a distinguished professor in the UAB School of Medicine, and he holds the Nancy E. Dunlap, M.D., Endowed Chair in Pulmonary Disease.
Reference
Genschmer KR, Russell DW, et al. Activated PMN Exosomes: Pathogenic Entities Causing Matrix Destruction and Disease in the Lung. Cell (2019) 176: 113-126. doi: 10.1016/j.cell.2018.12.002. PMID: 30633902.