Secreted RNAs leave the intracellular environment by associating with diverse vesicular and protein components. Secreted vesicles are heterogeneous and follow various routes of egress from the cell (1). Subclasses of such vesicles contain distinct cell surface proteins (2). In order to fully understand the diversity of vesicles that contain RNA, it is necessary to analyze and sort vesicle populations (3). One way to do this is by flow sorting such vesicles based on the presence of distinct vesicular surface proteins.
The ability to perform flow cytometric analysis and sorting of exosomes has been an ongoing area of controversy due to the small size of exosomes, which range in size from 40-130nm, near or below the diffraction limit of light. Nevertheless, a variety of groups have used this technique to analyze different subsets of small vesicles successfully (4-14), including proteomic analyses (4, 15-17). The efficacy of these flow-sorting experiments has been cross-validated by a variety of means, including western blots and co-localization of coincidently expressed factors. Fluorescence-Activated Vesicle Sorting (FAVS) uses light scattering properties of vesicles to analyze and sort individual exosomes using fluorescent labels. (See a previous blog on FAVS here.)
In the paper, “Identification and Characterization of EGF Receptor in Individual Exosomes by Fluorescence-Activated Vesicle Sorting (FAVS)”, published in the Journal of Extracellular Vesicles (JEV), Higginbotham and colleagues have used FAVS to analyze exosomal subsets that express varying amounts of EGFR in different cell-culture and in vivo contexts. This was done using DiFi cells, a human colorectal cancer (CRC) cell line, and A431, an epidermoid cancer cell line, which express approximately 5×106 and 2.5×106 EGFRs per cell, respectively (18, 19). The FAVS results showed that DiFi exosomes contain far more EGFR than do A431 exosomes, far exceeding the two-fold difference in EGFR levels present in these cell lines. Furthermore, using an antibody that recognizes an active form of EGFR, mAb806 (20-22), the amount of active EGFR was also found to be dramatically higher in DiFi exosomes than in A431 exosomes.
FAVS was also used to sort EGFR/CD9 double-positive and double-negative exosome populations, allowing enrichment of both subsets by post-sort analysis as well as western blot validation of the sorted exosomes (see Figure). Using human-specific reagents, FAVS was able to detect DiFi exosomes in the plasma of mice bearing DiFi xenografts. FAVS was also used to demonstrate that EGFR and one of its ligands, amphiregulin (AREG) are present in the plasma of normal individuals.
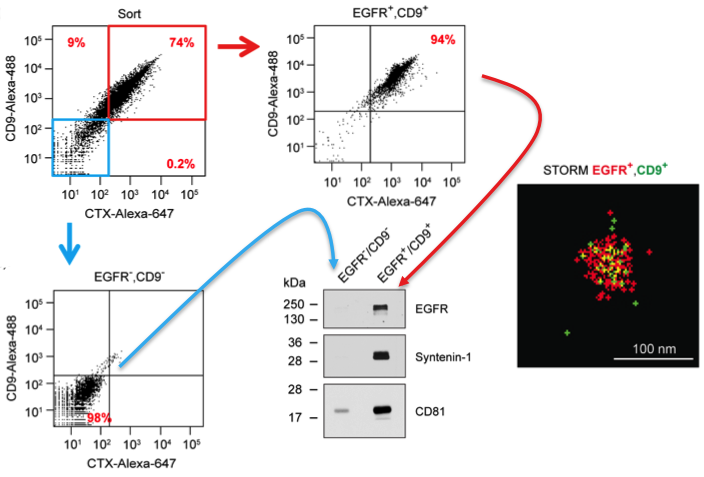
Results from the JEV paper derived from Fig 2. DiFi exosomes were flow sorted using antibodies to EGFR and CD9. Sorted purified double-negative vesicles (blue box/arrow) and double-positive vesicles (red box/arrow) were probed by western blot for markers as shown. These results validate the flow sorting enrichment of these different classes of vesicles. Also shown is a STORM image of an individual flow sorted double-positive vesicle.
This work joins flow-sorting work done by other labs using somewhat different techniques (6-14) and has implications for similar kinds of work done by other members of this consortium (23-25). Common to all these techniques was the use of lipid and/or specific extracellular vesicle markers to identify classes of secreted vesicles. Unlike FAVS, many sorting methods trigger vesicular events based on fluorescence rather than scatter. In all of these cases, analysis of secreted vesicle populations was performed. In some cases vesicle sorting was also achieved.
Thus, FAVS appears to be a promising technique to identify and purify distinct subsets of exosomes for discovery studies. It also holds promise for the detection of biomarkers in disease states including subsets of associated secreted RNAs.
1. Yang JM, Gould SJ. The cis-acting signals that target proteins to exosomes and microvesicles. Biochem Soc Trans (2013) 41:277-82. PMID 23356297.
2. Kowal J, et al. Proteomic comparison defines novel markers to characterize heterogeneous populations of extracellular vesicle subtypes. Proc Natl Acad Sci USA (2016) 113:E968-77. PMID 26858453.
3. Lunavat TR, et al. Small RNA deep sequencing discriminates subsets of extracellular vesicles released by melanoma cells–Evidence of unique microRNA cargos. RNA Biol (2015) 12:810-23. PMID 26176991.
4. Cao Z, et al. Use of fluorescence-activated vesicle sorting for isolation of Naked2-associated, basolaterally targeted exocytic vesicles for proteomics analysis. Mol Cell Proteomics (2008) 7:1651-67. PMID 18504258.
5. Higginbotham JN, et al. Amphiregulin exosomes increase cancer cell invasion. Curr Biol (2011) 21:779-86. PMID 21514161.
6. Poncelet P, et al. Standardized counting of circulating platelet microparticles using currently available flow cytometers and scatter-based triggering: Forward or side scatter? Cytometry A (2016) 89:148-58. PMID 25963580.
7. Erdbrugger U, Lannigan J. Analytical challenges of extracellular vesicle detection: A comparison of different techniques. Cytometry A (2016) 89:123-34. PMID 26651033.
8. Chandler WL, Yeung W, Tait JF. A new microparticle size calibration standard for use in measuring smaller microparticles using a new flow cytometer. J Thromb Haemost (2011) 9:1216-24. PMID 21481178.
9. van der Pol E, et al. Single vs. swarm detection of microparticles and exosomes by flow cytometry. J Thromb Haemost (2012) 10:919-30. PMID 22394434.
10. Arraud N, Gounou C, Turpin D, Brisson AR. Fluorescence triggering: A general strategy for enumerating and phenotyping extracellular vesicles by flow cytometry. Cytometry A (2016) 89:184-95. PMID 25857288.
11. Pospichalova V, et al. Simplified protocol for flow cytometry analysis of fluorescently labeled exosomes and microvesicles using dedicated flow cytometer. J Extracell Vesicles (2015) 4:25530. PMID 25833224.
12. Nolte-‘t Hoen EN, et al. Quantitative and qualitative flow cytometric analysis of nanosized cell-derived membrane vesicles. Nanomedicine (2012) 8:712-20. PMID 22024193.
13. van der Vlist EJ, et al. Fluorescent labeling of nano-sized vesicles released by cells and subsequent quantitative and qualitative analysis by high-resolution flow cytometry. Nat Protoc (2012) 7:1311-26. PMID 22722367.
14. Groot Kormelink T, et al. Prerequisites for the analysis and sorting of extracellular vesicle subpopulations by high-resolution flow cytometry. Cytometry A (2016) 89:135-47. PMID 25688721.
15. Demory Beckler M, et al. Proteomic analysis of exosomes from mutant KRAS colon cancer cells identifies intercellular transfer of mutant KRAS. Mol Cell Proteomics (2013) 12:343-55. PMID 23161513.
16. McConnell RE, et al. The enterocyte microvillus is a vesicle-generating organelle. J Cell Biol (2009) 185:1285-98. PMID 19564407.
17. Shifrin DA, et al. Enterocyte microvillus-derived vesicles detoxify bacterial products and regulate epithelial-microbial interactions. Curr Biol (2012) 22:627-31. PMID 22386311.
18. Gross ME, et al. Cellular growth response to epidermal growth factor in colon carcinoma cells with an amplified epidermal growth factor receptor derived from a familial adenomatous polyposis patient. Cancer Res (1991) 51:1452-9. PMID 1847663.
19. Kawamoto T, et al. Growth stimulation of A431 cells by epidermal growth factor: identification of high-affinity receptors for epidermal growth factor by an anti-receptor monoclonal antibody. Proc Natl Acad Sci USA (1983) 80:1337-41. PMID 6298788.
20. Walker F, et al. Ligand binding induces a conformational change in epidermal growth factor receptor dimers. Growth Factors (2012) 30:394-409. PMID 23163584.
21. Gan HK, et al. Targeting of a conformationally exposed, tumor-specific epitope of EGFR as a strategy for cancer therapy. Cancer Res (2012) 72:2924-30. PMID 22659454.
22. Reilly EB, et al. Characterization of ABT-806, a Humanized Tumor-Specific Anti-EGFR Monoclonal Antibody. Mol Cancer Ther (2015) 14:1141-51. PMID 25731184.
23. Stoner SA, et al. High sensitivity flow cytometry of membrane vesicles. Cytometry A (2016) 89:196-206. PMID 26484737.
24. Nolan JP. Flow Cytometry of Extracellular Vesicles: Potential, Pitfalls, and Prospects. Curr Protoc Cytom (2015) 73:13.4.1-6. PMID 26132176.
25. Danielson KM, et al. Diurnal Variations of Circulating Extracellular Vesicles Measured by Nano Flow Cytometry. PLoS ONE (2016) 11:e0144678. PMID 26745887.
Protocols for studying extracellular vesicles and extracellular RNA
Extracellular RNA in the Immune System symposium ignites interest at Immunology 2016
There are no comments.