The secretion of biomolecules into the extracellular milieu is a common phenomenon in biology. Similar to mammalian cells which secrete and use extracellular vesicles to communicate between cells using the contained biomolecules; bacteria, e.g. Escherichia coli, also secrete outer membrane vesicles (OMVs (*); Table 1) containing distinct biomolecules which are thought to be involved in intra-species communication, in inter-kingdom exchanges and pathogenicity (1–5). To date, such exported molecules such as small molecules, DNA, peptides and proteins, have been well-studied in bacteria (2,5,6). The bacterial extracellular RNA complement has only been very recently characterized. More specifically, a very recent report has found that Prochlorococcus, the numerically dominant marine cyanobacterium, continuously releases lipid vesicles containing proteins, DNA and RNA into its extracellular milieu (7). In total, 89% of the genome of Prochlorococcus was represented at least once in the RNA-associated libraries from the OMV fraction (7).
Using a combination of physical characterization and RNA sequencing, our group has recently analyzed the extracellular RNA complement of both OMV-associated and OMV-free RNA of the enteric Gram-negative model bacterium Escherichia coli K-12 substrain MG1655, and we have compared it to its intracellular RNA complement (8). Our results demonstrate that when MG1655 is cultured in LB, rich bacterial media, a large part of the extracellular RNA complement is in the size range between 15 and 40 nucleotides (Figure 1) and is derived from specific intracellular RNA species.
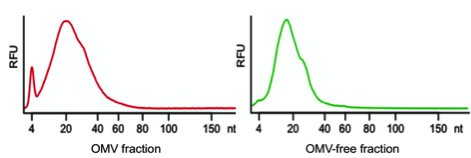
Figure 1: Size distribution of extracellular RNA released by Escherichia coli K-12. In red the size distribution of RNA associated with OMVs is represented and in green RNA extracted from the OMV-free bacterial supernatant is shown. (RFU: relative fluorescence unit).
In addition, we demonstrated that RNA is specifically associated with OMVs (Figure 2) and that the relative abundances of RNA biotypes in the intracellular, OMV and OMV-free fractions are quite distinct (Figure 3).
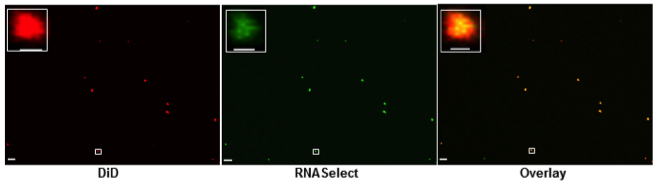
Figure 2: Confocal microscopy analysis of OMVs, stained with lipid tracer dye, DiD (red) and RNA specific dye, SYTO RNASelect (green). The area highlighted within the white rectangular box is magnified in the inset. The scale bar is equivalent to 5 µm in the main images and equivalent to 500 nm in the magnified images. Each individual colour dot in the images likely represents the aggregation of several OMVs, as the typical sizes of OMVs are below the limit of resolution of confocal microscopy.
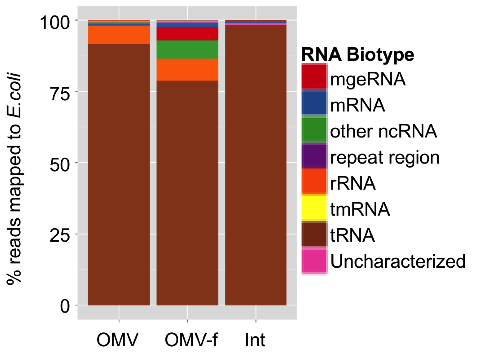
Figure 3: Annotated profile of RNA extracted from OMVs obtained from Escherichia coli cultures (OMV), of RNA extracted from OMV-depleted Escherichia coli culture supernatant (OMV-f) and RNA extracted from the bacterial cells themselves (Int).
Apart from rRNA fragments, a significant portion of the extracellular RNA complement is composed of specific cleavage products of functionally important structural, non-coding RNAs, including tRNAs, 4.5S RNA, 6S RNA and tmRNA (transfer-messenger RNA: bacterial RNA with dual tRNA-like and mRNA-like properties).
The function of these exported small RNAs is still unknown, but it is important to note that OMVs can be taken up by human host cells. Thus, they may even contribute to pathogenesis, as demonstrated for OMVs derived from Helicobacter pylori, which are taken up by gastric epithelial cells (9). In this context, OMVs have been found to enhance the carcinogenic potential of this specific bacterium (10).
As the number of microorganisms living in and on the human body outnumbers the total number of human cells by at least an order of magnitude and knowing that the vesicular secretion by bacteria is a commonly observed phenomenon (11–14), it is not surprising that first reports have shown that OMVs can influence human health and specific disease states (Table 1). OMVs released by gastrointestinal pathogens can be harmful or even lethal, as OMVs derived from pathogenic bacteria transport diverse virulence factors to the host cells, enabling them to modulate host defense and response in order to assure their survival and replication (11,13,15,16) (Table 1). On the other hand, it has only recently become appreciated that OMV-induced signaling by commensal bacteria is of utmost importance for the host (5). A recent study shows that polysaccharide A enriched OMVs derived from a human commensal (Bacteroides fragilis) mediate host immune regulation and prevent colitis in a mouse model (5).
Our results, described in Ghosal et al., suggest a selective export of specific RNA biotypes by Escherichia coli (8). For example, specific fragments of specific tRNAs are found in the OMV, and are different from those found outside the OMVs. In addition, our unpublished data suggests that Salmonella enterica serovar Typhimurium also secretes small RNA-enriched OMVs (personal communication). These initial results seem promising, but require more detailed and mechanistic studies in order to ascertain if bacterial secreted small RNAs do play a role in intercellular communication, and what those roles may be. This is entirely analogous to findings in eukaryotes, where small RNA can be delivered via vesicles to their target cell and trigger a response (17–19). Indeed, a previous study in bacteria seems to underscore this hypothesis as it demonstrates that extracellular RNA secreted by Listeria spp. is a key component for the host developing an immune response against the bacterial infection (20). Moreover, small extracellular RNA fragments of Mycobacterium tuberculosis (comprised primarily of tRNA and rRNA fragments), induce early apoptosis in human monocytes (21). Whether these molecules are delivered to their target via OMVs still needs to be conclusively established.
Understanding the role of bacteria-derived exogenous RNA in host-microbe interactions, in pathogenesis as well as in mutualism, will elucidate new mechanisms and perhaps allow the identification of new drug targets and/or the development of RNA-based vaccines. Further investigations in the field of extracellular bacterial RNAs are clearly needed to shed light on their potential role as mediators of microbe-microbe and host-microbe intercellular communication, and on the specific mechanisms of these effects. This will be an exciting advance that could provide entirely new approaches to human disease therapy and prevention.
*: OMV definition and functions
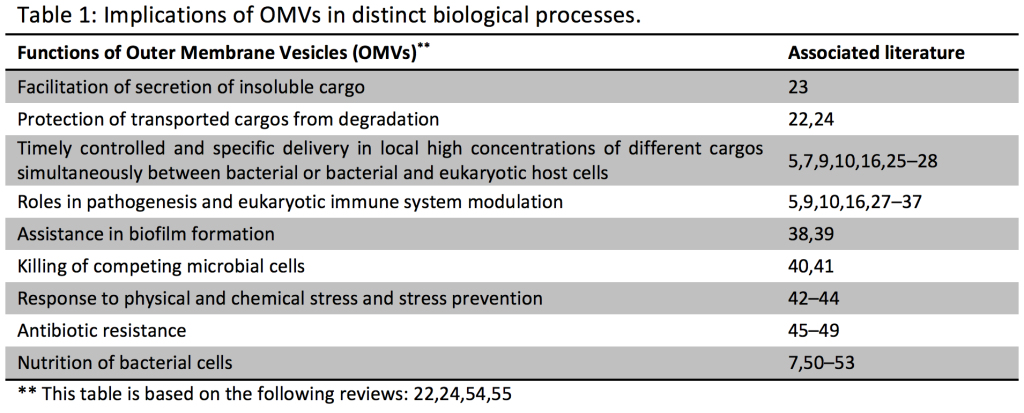
Outer membrane vesicles (OMV) are produced by all Gram-negative bacteria. They are thought to form when buds from the outer membrane (OM) of cells encapsulate periplasmic material and pinch off from the OM to form spheroid particles with a size range of 10-300 nm in diameter. By the inclusion of cargos (small molecules, peptides and proteins, DNA, RNA) into OMVs, the cargos may benefit from : protection from degradation, long-distance delivery, specificity in host-cell targeting, evasion of the hosts immune response and coordinated secretion with other bacterial effectors (22).
References
1. Tseng T-T, Tyler BM, Setubal JC. Protein secretion systems in bacterial-host associations, and their description in the Gene Ontology. BMC Microbiol. 2009;9 Suppl 1:S2.
2. Molloy S. Setting the threshold. Nat. Rev. Microbiol. 2010;8(June):
3. Lee VT, Schneewind O. Protein secretion and the pathogenesis of bacterial infections. Genes Dev. 2001;15(617):1725–1752.
4. Hughes DT, Sperandio V. Inter-kingdom signalling: communication between bacteria and their hosts. Nat. Rev. Microbiol. 2008;6(2):111–20.
5. Shen Y, Giardino Torchia ML, Lawson GW, et al. Outer membrane vesicles of a human commensal mediate immune regulation and disease protection. Cell Host Microbe. 2012;12(4):509–20.
6. Waters CM, Bassler BL. Quorum sensing: cell-to-cell communication in bacteria. Annu. Rev. Cell Dev. Biol. 2005;21:319–46.
7. Biller SJ, Schubotz F, Roggensack SE, et al. Bacterial Vesicles in Marine Ecosystems. Science (80-. ). 2014;998(January):183–187.
8. Ghosal A, Upadhyaya BB, Fritz JV., et al. The extracellular RNA complement of Escherichia coli. Microbiologyopen. 2015;n/a–n/a.
9. Parker H, Chitcholtan K, Hampton MB, Keenan JI. Uptake of Helicobacter pylori outer membrane vesicles by gastric epithelial cells. Infect. Immun. 2010;78(12):5054–61.
10. Chitcholtan K, Hampton MB, Keenan JI. Outer membrane vesicles enhance the carcinogenic potential of Helicobacter pylori. Carcinogenesis. 2008;29(12):2400–5.
11. Kuehn MJ, Kesty NC. Bacterial outer membrane vesicles and the host-pathogen interaction. Genes Dev. 2005;19(22):2645–55.
12. Wai SN, Lindmark B, Söderblom T, et al. Vesicle-mediated export and assembly of pore-forming oligomers of the enterobacterial ClyA cytotoxin. Cell. 2003;115(1):25–35.
13. Park K-S, Choi K-H, Kim Y-S, et al. Outer membrane vesicles derived from Escherichia coli induce systemic inflammatory response syndrome. PLoS One. 2010;5(6):e11334.
14. Thay B, Wai SN, Oscarsson J. Staphylococcus aureus α-toxin-dependent induction of host cell death by membrane-derived vesicles. PLoS One. 2013;8(1):e54661.
15. Mashburn-Warren LM, Whiteley M. Special delivery: vesicle trafficking in prokaryotes. Mol. Microbiol. 2006;61(4):839–46.
16. Bomberger JM, Maceachran DP, Coutermarsh B a, et al. Long-distance delivery of bacterial virulence factors by Pseudomonas aeruginosa outer membrane vesicles. PLoS Pathog.
2009;5(4):e1000382.
17. Li CCY, Eaton SA, Young PE, et al. Glioma microvesicles carry selectively packaged coding and non-coding RNAs which alter gene expression in recipient cells. RNA Biol. 2013;10(August):1333–1344.
18. Buck AH, Coakley G, Simbari F, et al. Exosomes secreted by nematode parasites transfer small RNAs to mammalian cells and modulate innate immunity. Nat. Commun. 2014;5:5488.
19. Khalyfa A, Gozal D. Exosomal miRNAs as potential biomarkers of cardiovascular risk in children. J. Transl. Med. 2014;12(1):162.
20. Abdullah Z, Schlee M, Roth S, et al. RIG-I detects infection with live Listeria by sensing secreted bacterial nucleic acids. EMBO J. 2012;31(21):4153–64.
21. Obregón-Henao A, Duque-Correa M a, Rojas M, et al. Stable extracellular RNA fragments of Mycobacterium tuberculosis induce early apoptosis in human monocytes via a caspase-8 dependent mechanism. PLoS One. 2012;7(1):e29970.
22. Bonnington KE, Kuehn MJ. Protein selection and export via outer membrane vesicles. Biochim. Biophys. Acta. 2014;1843(8):1612–9.
23. Greniert D, Mayrand D. Produced by. Infect. Immun. 1987;55(1):111–117.
24. Kulkarni HM, Jagannadham M V. Biogenesis and multifaceted roles of outer membrane vesicles from Gram-negative bacteria. Microbiology. 2014;160(Pt 10):2109–21.
25. Yaron S, Kolling GL, Simon L, Matthews KR. Vesicle-mediated transfer of virulence genes from Escherichia coli O157:H7 to other enteric bacteria. Appl. Environ. Microbiol.
2000;66(10):4414–20.
26. Fulsundar S, Harms K, Flaten GE, et al. Gene transfer potential of outer membrane vesicles of Acinetobacter baylyi and effects of stress on vesiculation. Appl. Environ. Microbiol. 2014;80(11):3469–83.
27. Furuta N, Tsuda K, Omori H, et al. Porphyromonas gingivalis outer membrane vesicles enter human epithelial cells via an endocytic pathway and are sorted to lysosomal compartments. Infect. Immun. 2009;77(10):4187–96.
28. Furuta N, Takeuchi H, Amano A. Entry of Porphyromonas gingivalis outer membrane vesicles into epithelial cells causes cellular functional impairment. Infect. Immun. 2009;77(11):4761–70.
29. Horstman AL, Kuehn MJ. Bacterial surface association of heat-labile enterotoxin through lipopolysaccharide after secretion via the general secretory pathway. J. Biol. Chem. 2002;277(36):32538–45.
30. Kesty NC, Mason KM, Reedy M, Miller SE, Kuehn MJ. Enterotoxigenic Escherichia coli vesicles target toxin delivery into mammalian cells. EMBO J. 2004;23(23):4538–49.
31. Karavolos MH, Bulmer DM, Spencer H, et al. Salmonella Typhi sense host neuroendocrine stress hormones and release the toxin haemolysin E. EMBO Rep. 2011;12(3):252–8.
32. Schild S, Nelson EJ, Camilli A. Immunization with Vibrio cholerae outer membrane vesicles induces protective immunity in mice. Infect. Immun. 2008;76(10):4554–63.
33. Ellis TN, Leiman SA, Kuehn MJ. Naturally produced outer membrane vesicles from Pseudomonas aeruginosa elicit a potent innate immune response via combined sensing of both lipopolysaccharide and protein components. Infect. Immun. 2010;78(9):3822–31.
34. Vidakovics ML a P, Jendholm J, Mörgelin M, et al. B cell activation by outer membrane vesicles–a novel virulence mechanism. PLoS Pathog. 2010;6(1):e1000724.
35. Nakao R, Hasegawa H, Ochiai K, et al. Outer membrane vesicles of Porphyromonas gingivalis elicit a mucosal immune response. PLoS One. 2011;6(10):e26163.
36. Elmi A, Watson E, Sandu P, et al. Campylobacter jejuni outer membrane vesicles play an important role in bacterial interactions with human intestinal epithelial cells. Infect. Immun. 2012;80(12):4089–98.
37. Pollak CN, Delpino MV, Fossati CA, Baldi PC. Outer membrane vesicles from Brucella abortus promote bacterial internalization by human monocytes and modulate their innate immune response. PLoS One. 2012;7(11):e50214.
38. Beveridge TJ, Makin SA, Kadurugamuwa JL, Li Z. Interactions between biofilms and the environment. FEMS Microbiol. Rev. 1997;20(3-4):291–303.
39. Baumgarten T, Sperling S, Seifert J, et al. Membrane vesicle formation as a multiple-stress response mechanism enhances Pseudomonas putida DOT-T1E cell surface hydrophobicity and biofilm formation. Appl. Environ. Microbiol. 2012;78(17):6217–24.
40. Li Z, Clarke AJ, Beveridge TJ. Gram-negative bacteria produce membrane vesicles which are capable of killing other bacteria. J. Bacteriol. 1998;180(20):5478–83.
41. Vasilyeva N V, Tsfasman IM, Suzina NE, Stepnaya OA, Kulaev IS. Secretion of bacteriolytic endopeptidase L5 of Lysobacter sp. XL1 into the medium by means of outer membrane vesicles. FEBS J. 2008;275(15):3827–35.
42. McBroom AJ, Kuehn MJ. Release of outer membrane vesicles by Gram-negative bacteria is a novel envelope stress response. Mol. Microbiol. 2007;63(2):545–58.
43. Manning AJ, Kuehn MJ. Contribution of bacterial outer membrane vesicles to innate bacterial defense. BMC Microbiol. 2011;11:258.
44. Loeb MR, Kilner J. Release of a special fraction of the outer membrane from both growing and phage T4-infected Escherichia coli B. Biochim. Biophys. Acta. 1978;514(1):117–27.
45. Ciofu O, Beveridge TJ, Kadurugamuwa J, Walther-Rasmussen J, Høiby N. Chromosomal beta-lactamase is packaged into membrane vesicles and secreted from Pseudomonas aeruginosa. J. Antimicrob. Chemother. 2000;45(1):9–13.
46. Schaar V, Nordström T, Mörgelin M, Riesbeck K. Moraxella catarrhalis outer membrane vesicles carry β-lactamase and promote survival of Streptococcus pneumoniae and Haemophilus influenzae by inactivating amoxicillin. Antimicrob. Agents Chemother. 2011;55(8):3845–53.
47. Schaar V, Paulsson M, Mörgelin M, Riesbeck K. Outer membrane vesicles shield Moraxella catarrhalis β-lactamase from neutralization by serum IgG. J. Antimicrob. Chemother. 2013;68(3):593–600.
48. Schaar V, Uddbäck I, Nordström T, Riesbeck K. Group A streptococci are protected from amoxicillin-mediated killing by vesicles containing β-lactamase derived from Haemophilus influenzae. J. Antimicrob. Chemother. 2014;69(1):117–20.
49. Mashburn LM, Whiteley M. Membrane vesicles traffic signals and facilitate group activities in a prokaryote. Nature. 2005;437(7057):422–5.
50. Kulp A, Kuehn MJ. Biological functions and biogenesis of secreted bacterial outer membrane vesicles. Annu. Rev. Microbiol. 2010;64:163–84.
51. Lee E-Y, Bang JY, Park GW, et al. Global proteomic profiling of native outer membrane vesicles derived from Escherichia coli. Proteomics. 2007;7(17):3143–53.
52. Elhenawy W, Debelyy MO, Feldman MF. Preferential packing of acidic glycosidases and proteases into Bacteroides outer membrane vesicles. MBio. 2014;5(2):e00909–14.
53. Rakoff-Nahoum S, Coyne MJ, Comstock LE. An ecological network of polysaccharide utilization among human intestinal symbionts. Curr. Biol. 2014;24(1):40–9.
54. Avila-Calderón ED, Araiza-Villanueva MG, Cancino-Diaz JC, et al. Roles of bacterial membrane vesicles. Arch. Microbiol. 2015;197(1):1–10.
55. Haurat MF, Elhenawy W, Feldman MF. Prokaryotic membrane vesicles: new insights on biogenesis and biological roles. Biol. Chem. 2015;396(2):95–109.